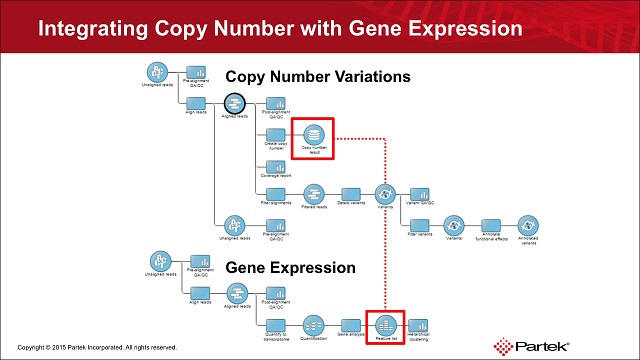
Detecting Copy Number Variations and Integrating with Gene Expression
Copy number variations (CNV) are a common form of structural variation, accounting for approximately 10% of human genomic DNA. Although CNVs receive less attention than do single nucleotide variations (SNVs), they are a risk or a protective factor for various diseases and account for differences between individuals. A possible consequence of CNVs is an increase or decrease of expression levels of affected genes, a phenomenon that can be studied by combining DNA-Seq with RNA-Seq data. In this video we will teach you how to detect copy number variations in your NGS data and how to integrate it into your gene expression study.
Learn how to
- Analyzing whole genome sequencing data
- Detecting amplifications and deletions
- Interpreting the results through pathway analysis
- Integrating copy number with gene expression data
Register to Watch
Share this post:
