Integration of Copy Number and Methylation Data from the Illumina® Infinium HumanMethylation450 BeadChip Array
Integration of genomic data and epigenomic data as a means for understanding complex genomic regulatory mechanisms is gaining in popularity. In this webinar, we will show you how to use Partek® Genomics Suite® to identify copy number regions from the Illumina® Infinium HumanMethylation450 BeadChip array and merge the results with differentially methylated regions detected using the same arrays.
Learn how to:
- Import Illumina .idat files and normalize the data
- Detect differentially methylated regions
- Remove probes with SNP in the vicinity of the query site
- Visualize the methylation signature
- Create a list of genes regulated by differential methylation
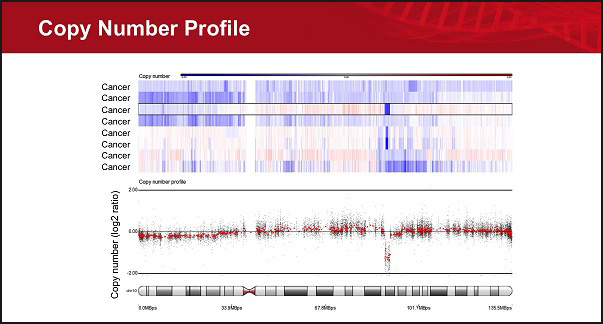
- Identify copy number regions
- Find copy number regions shared across study samples
- Integrate the copy number data with methylation data
Register to Watch
Share this post:
